Chemical fingerprints are used for both similarity and substructure searching. When used for similarity, a score accounts for features shared and different between compounds. For substructure searching, the fingerprint provides a prescreen of potential hits by enforcing that every feature encoded in the query fingerprint must be present in the reference. If a single feature is found in the query but not in the reference can it safely be discarded?
Chemical fingerprints are used for both similarity and substructure searching. When used for similarity, a score accounts for features shared and different between compounds. For substructure searching, the fingerprint provides a prescreen of potential hits by enforcing that every feature encoded in the query fingerprint must be present in the reference. If a single feature is found in the query but not in the reference can it safely be discarded?
Common types of fingerprints include: substructure keys (MACCS, CACTVS), path (Daylight), circular (ECFP, Morgan), tree, and n-gram (LINGOS, IBM). The fingerprint examples described below are often documented as being similarity fingerprints or “optimised for similarity” but it isn’t always stated that their use should be avoid for substructure screening. A fingerprint intended for similarity will often screen out results from a substructure search that do actually match (false negatives).
Connectivity
Circular and n-gram fingerprints inherently can not be used for substructure filtering as they capture the absence as well as the presence of neighbours [1].
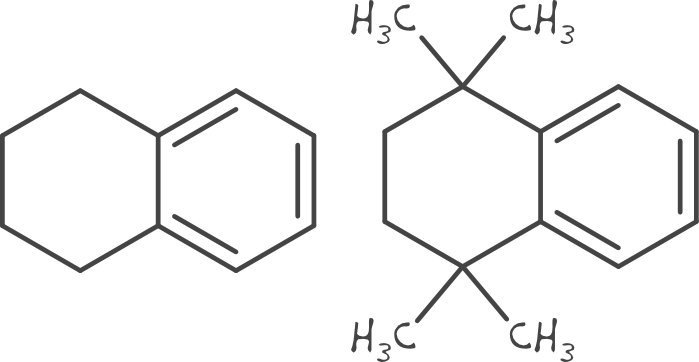
As is seen with the circular fingerprint, the number of neighbours (degree) is not invariant between the query and reference. The degree of the reference atoms must be equal or more to that of the query. The connectivity/degree can therefore not be encoded in the other types of fingerprints.
Hydrogen count
Similar to connectivity, the hydrogen count may be less than or greater than the query. The MACCS 166 substructure keys (as used in the open source toolkits) were reoptimised for similarity[2,3]. As some keys match hydrogen counts, they should not be used as a substructure fingerprint:
In the compounds above, the MACCS keys 118 (‘[#6H2]([#6H2]*)*’>1) and 129 (‘[#6H2](~*~*~[#6H2]~*)~*’) are found in the query (left) but not the reference (right). The CACTVS substructure keys also match hydrogens (e.g. bit 329, 335) and have the same property. As with MACCS, the documentation states that the CACTVS keys are intended for similarity.
Hybridisation
Attempting to encode hybridisation is also problematic, consider the following query and target.
The left is not considered a substructure of the right with the CDK’s hybridisiation fingerprinter as an sp2 carbon in the query is sp1 in the reference.
Rings
Care should also be taken with ring size, in particular the smallest ring size of an atom or bond is not invariant.
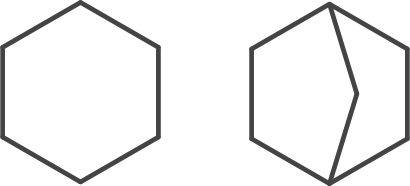
This behaviour is observed with the CDK’s ShortestPath fingerprint where the query (left) has atoms in a smallest ring of size six but the reference (right) has atoms in either smaller ring of size five. More subtle issues are found when using the non-unique SSSR [4]. Some effects of the use of (E)SSSR are observed in the CACTVS substructure keys (intended for similarity as stated in the manual).
For these two PubChem Compound entries (CID 135973, CID 9249) the query (left) encodes a four membered ring while the reference (right) does not.
It is possible to encode and match the degree and hydrogen count in a fingerprint just not as a single feature. Encoding the degree in the feature or layering properties (ala RDKit Fingerprint) can be done safely but is redundant and leads to a denser fingerprint. Ring size information can also be encoded, rather than encoding smallest rings, all ring sizes (up to some length) need to be encoded.
Take home message
Different fingerprints exist for different purposes and surprisingly few are truly suitable for substructure filtering. Path and tree fingerprints are generally okay but caution must be taken to ensure variant properties are not encoded. The keen eyed may notice there is no mentioned of issues with aromaticity in fingerprints; there are unfortunately too many to list in a single post.
- http://pubs.acs.org/doi/abs/10.1021/ci100050t
- http://pubs.acs.org/doi/abs/10.1021/ci010132r
- http://www.dalkescientific.com/writings/diary/archive/2011/01/20/implementing_cactvs_keys.html
- http://docs.eyesopen.com/toolkits/oechem/cplusplus/ring.html#smallest-set-of-smallest-rings-sssr-considered-harmful
Image credit: CPOA