In 2012, 2014 and 2017 Daniel Lowe (while at NextMove Software) released a large collection of reactions extract from USPTO patent applications as CC-Zero. We have made updates (currently quarterly) for customers of our Pistachio query tool and extended the reaction data to include USPTO Sketches and European patents.
The next version of Pistachio will include data from WIPO PCT documents and include several enhancements to content and representation. Highlight these in a blog post made more sense than the usual release notes.
Number of Reactions
The next release will contain >13,118,970 reactions from the following sources:
| Source | Count | Latest Extraction |
| USPTO Grant Text | 3,290,056 | 2021-03-16 |
| USPTO Appl. Text | 3,595,510 | 2021-03-18 |
| WIPO PCT Text | 1,484,646 | 2021-03-18 |
| EPO Grant Text | 1,060,397 | 2021-03-17 |
| EPO Appl. Text | 696,578 | 2021-03-17 |
| USPTO Grant Sketch | 1,186,924 | 2021-03-16 |
| USPTO Appl. Sketch | 1,804,859 | 2021-03-18 |
Pistachio covers 1976-current day however 4,447,418 (33%) were published in the last five years and 8,166,128 (62%) in the last ten years.
The reaction data is document (or citation) centric, the same reaction text often occurs in an application and grant. It can also be published in multiple authorities (e.g. USPTO, EPO and WIPO). Often the description text is identical but not always, sometimes a product yield/quantity may be miss-typed or omitted. The number of unique reactions by RInChI is 4,212,894.
All reactions extracted from text are Atom-Atom Mapped either by NameRxn or if the reaction is unrecognised we fallback to Indigo. 9,383,607 (71.5%) are currently recognized by NameRxn.
What’s New?
Through improvements related to the WIPO PCT inclusion we observed a ~15% increase in recall for existing USPTO and EPO data. This is one of the many advantages of automated extraction over manually curated databases. Tweaks can be made and mistakes can fixed then applied in bulk over all the original source documents. Re-extraction currently takes a few days on a single machine (8 cores). Where miss-extraction mistakes are spotted we welcome the feedback and aim to resolve these where possible.
Embedded Heading Detection
One of the biggest challenges with handling WIPO PCT data is the English text is primarily OCRd. The submitted document quality can vary considerably which leads to a wide spectrum of related problems.
OCR is well known to have issues with the non-standard characters used in systematic chemical names. Fortunately the majority of issues are simple character transliterations and extra white space. These can be effectively handled with our spelling correction algorithms. Some very badly corrupted names are beyond all hope:

Part of the chemical name could not be OCRd and remains as an image
Another common issue with OCR is the detection of paragraph breaks and lack of title markup. Often chemical reaction descriptions use anaphoric references “title compound”, “desired product”, and “product from Step B” that need to be resolved. If the title is not found we can’t resolve it.
Paragraph breaks may be omitted:

There should be a paragraph break before “Intermediate 2:”
or in the wrong place (here splitting a chemical name):

The “Step H” compound has a paragraph break in the middle of the name.
To compensate for this, new algorithms were introduced to detect patterns of embedded headings where the break was missed by OCR. Existing inline heading (start of paragraph) detection was also improved.
These errors are not unique to WIPO data and occasionally occur in the USPTO documents too:

There should be a break before “Step-2:”
Patentscope (WIPO) recently announced a new OCR extraction framework that should improve paragraph detection and chemical formula recognition (from Feb 11th 2021). We have not yet found an improvement in the extraction recall rate.
Multi-Paragraph Reactions
In previous versions, the reaction step parsing started a new reaction on every new paragraph. This logic was tweaked to allow a reaction to span multiple paragraphs and handle the less reliable breaks in WIPO data. Regressions in USPTO extraction helped identify places in the reaction descriptions where a yielded product action was missed.
A side effect of this is we now sometimes extract cases where there was some unknown intermediate (A -> ?, ? -> B) as a single reaction. We are considering how to handle these.
Prefer Connected Representations
Chemical structures are generated from systematic names, line formulae, and dictionaries. Where possible we have updated the structure representations to favour connected representations:
[K]OC(=O)O[K] instead of [K+].[K+].[O-]C(=O)[O-] [Na][Cl] instead of [Na+].[Cl-] etc
Using CXSMILES fragment groups it is possible to keep the grouping of the counter-ions. The reaction components were also listed separately in the raw JSON files. Not all downstream tools can consume the CXSMILES representation and some users invented/adapted syntaxes to handle in their use cases e.g.
>[K+]+[K+]+[O-]C(=O)[O-]> >[K+]..[K+]..[O-]C(=O)[O-]>
The philosophy in preferring the connected component is that it is easier to split things apart then piece them back together (Humpty Dumpty). Note that not all counter-ions have been bonded due to undesirable valence representations (e.g. HATU, NH4Cl) and so the CXSMILES fragment groups remain a useful extension.
Representation of stereoisomer mixtures
I recently added the ability to OPSIN to capture racemic and relative stereo information in systematic names. In total around ~1% of reactions now have this information captured in CXSMILES. In a simple example we have an unknown mixture of enantiomers formed:
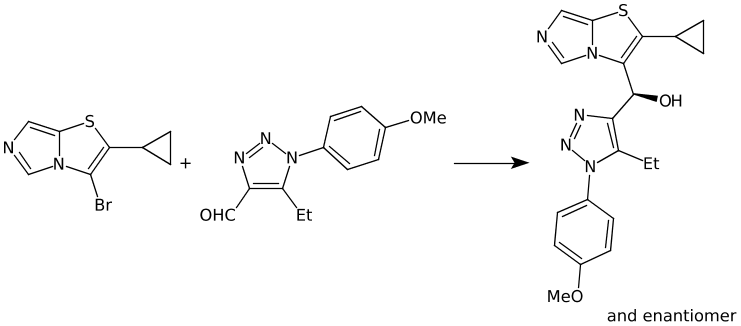
BrC1N2C=NC=C2SC=1C1CC1.C(C1N(C2C=CC(OC)=CC=2)N=NC=1C=O)C>>C1(C2SC3=CN=CN3C=2[C@H](C2N=NN(C3C=CC(OC)=CC=3)C=2CC)O)CC1 |&1:38| US20200405696A1_1398 Example 279
A more complex example is where one stereocenter configuration is known but the other is not:

ClC1C(CN2C(=O)N3[C@@H](C(N4CC[C@H](F)C4)=O)CN(C(OC(C)(C)C)=O)CC3=N2)=NC=CC=1C(F)(F)F>ClCCl.FC(F)(F)C(O)=O>ClC1C(CN2C(=O)N3[C@@H](C(N4CC[C@H](F)C4)=O)CNCC3=N2)=NC=CC=1C(F)(F)F |&1:8,55,a:13,60| US20200375986A1_0652 Example 4
Both of these cases could alternative be represented by simply removing the configuration from the racemic atom (and we may choose to normalise to that in future). The most important cases are when the relative configuration of two centres is known but that there is a mixture of enatiomers.

C(OC([C@H]1[C@H](C)CN(CC2C=CC=CC=2)C1)=O)C.CC(OC(OC(OC(C)(C)C)=O)=O)(C)C>C(O)C.[OH-].[OH-].[Pd+2]>C[C@H]1CN(C(OC(C)(C)C)=O)C[C@@H]1C(OCC)=O |f:3.4.5,&1:3,4,40,51| US20200369658A1_0530 Intermediate 2, Step b
NameRxn
Recent updates to NameRxn include limited support for some common non-balanced Functional Group Interconversion and Addition reactions, i.e, “Hydroxy to chloro” and “Amination”. This work also allowed a small performance boost. The total number of reaction types named is now 1,528 (from 1,297 previously), one source for additional reactions has been RXNO. NameRxn was not originally designed to provide Atom-Atom Maps; as that has become more of an interest we have made improvement to AAM where a functional group source was unknown.
Solvent Mixture Representations
More information about solvents and solvent mixtures is captured, for example: “5-chloro-3-((trimethylsilyl)ethynyl)pyrazin-2-amine (100 mg, 0.44 mmol) in THF (8 mL)” (US20200085822A1 Example 1 Step 6) the THF solvent is associated by reference from the reactant.
Finer grained details on component/volume fractions of solvent mixtures is also captured when described as “1M HCl Et2O”, “THF/MeOH (1 mL, 1:1)”
Sequence and Step Labels
Where found in the text we now include and attach the sequence and step labels to reactions. For example “Example 4, Step A”, “Compound 7, Step 2”. Pistachio will allow searching by the labels and resolving queries:

Azide-alkyne Huisgen cycloaddition (4.1.4)
Improved cross reference handling
Previously the cross-reference “Compound 1” was indexed and resolved just as “1”. We now use the “reference type” to disambiguate cases when there is both a “Compound 1” and “Intermediate 1”. The variety of recognised identifier values was also extended.
Summary
We have made several improvements to reaction extraction. These will be available in the new version of Pistachio that will be available at the start of next month.