While advanced software now exists for converting systematic chemical names to structures, the humble chemical line formula has for the most part avoided the limelight. For a long time LeadMine has been able to recognize chemical line formulae but we are now adding the ability to interpret them.
A simple example would be CHF3 which is 
Line formulae are interpreted from left-to-right, but in such a way that valency rules are respected e.g. the fluorine in the above example bonds to the C not the H.
More complicated examples include:
CF2=CF-O-CF2CF(CF3)O-CF2CF2-CH2OH [has an explicit double bond] {from US20020002258A1}
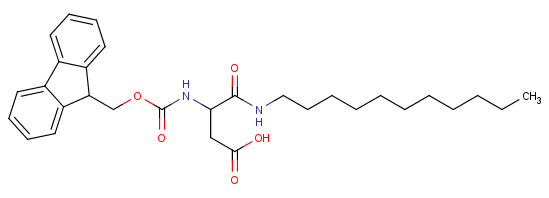
HO2C-CH2CH(NHFmoc)-CONH-(CH2)10CH3 [contains an abbreviated prefix: Fmoc] {from US20010038824A1}
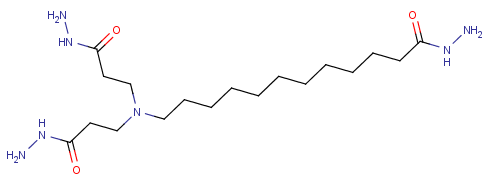
(NH2NHCOCH2CH2)2N(CH2)11CONHNH2 [has a repeated substituent, repeated infix and implicit double bonds for the carbonyls] {from US20050081961A1}
This is useful for pulling out reagents of chemical reactions and where the described compound is important in its own right.