A recurring task in medicinal chemistry is the optimization of R groups around a ring in order to improve biological activity. In a landmark paper in 1972, John Topliss (then at Schering) described a scheme for deciding what substituted phenyl to make next based on the relative potencies of the compounds made so far. This has become known as the Topliss Tree.
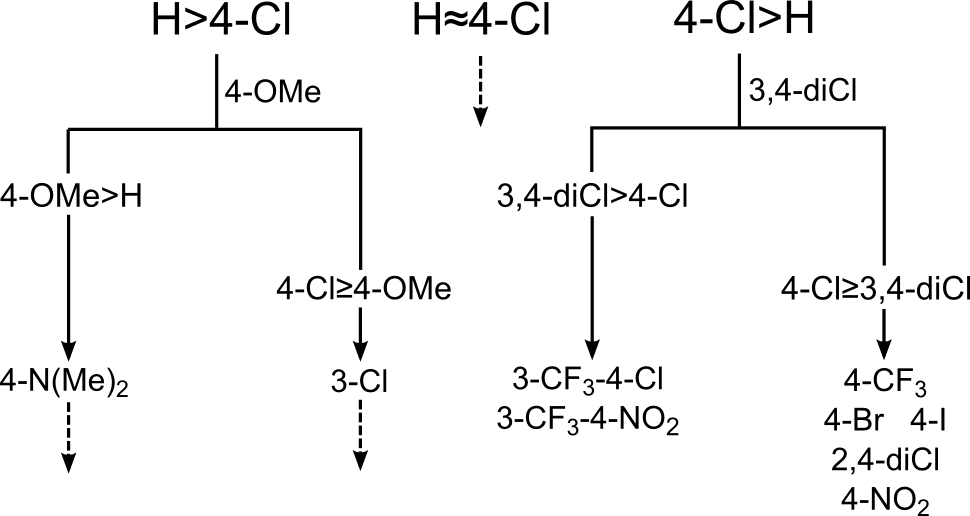
To begin with, Topliss suggested making the unsubstituted phenyl and the 4-chlorophenyl. Depending on their relative potencies, he went on to suggest either 4-methoxyphenyl or 3,4-dichlorophenyl. Based on the relative potency of that, the tree continued downwards. An abbreviated version of the tree is shown below.
So how did Topliss come up with his suggestions? He based it all on the Hansch parameters (σ and π) for different phenyl substituents, and the inferred relationship (either positive or negative) between the Hansch parameters and the potency. (This is less complicated than it sounds – it’s like identifying that potency is increasing with electron-withdrawing strength and so let’s try a stronger EWG.)
Today we have access to large amounts of experimental data on potency orders (for example, in ChEMBL), and so we can check whether that data agrees with the suggestions of the Topliss Tree. Back in March, we published a paper describing a method that uses a database of experimental results to suggest what R group would increase activity based on observed activity order. This is exactly the scenario described by Topliss and so we can come at the same question from a compeletely different angle. I briefly addressed this question in the paper, but I returned to it in more detail (and with more experimental data) for a talk I gave to MEDI at the recent ACS meeting in San Francisco: Revising the Topliss decision tree based on 30 years of medicinal chemistry literature [PDF]
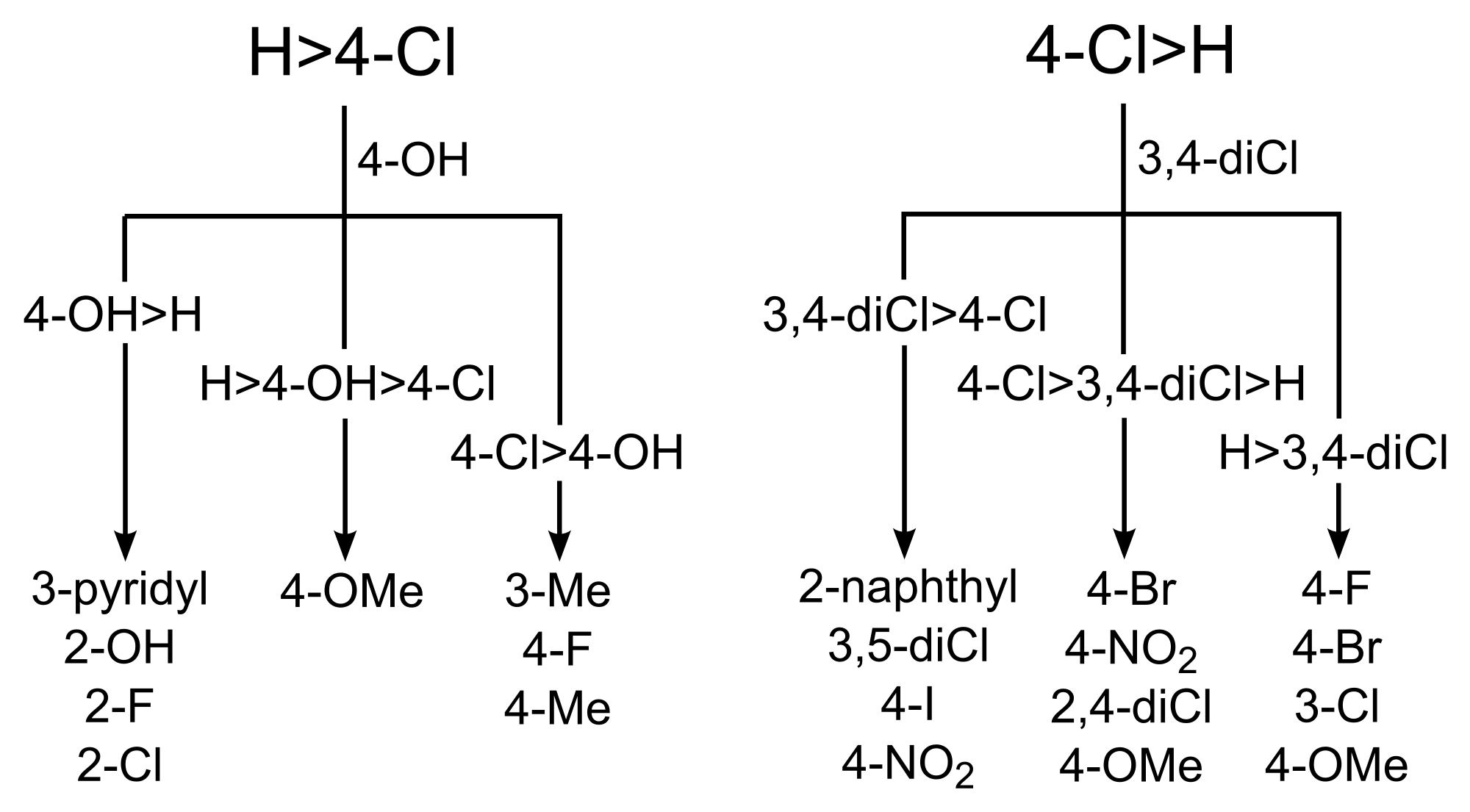
The conclusions were that in the main the data in ChEMBL agreed with Topliss. However, particular points of disagreement were the suggestion of 4-OMe instead of 4-OH (if H > 4-Cl), and the suggestion of 4-CF3 (if 4-Cl > 3,4-diCl). This then raises the question, what would we recommend instead?…enter the Matsy Tree:
Since we can generate a similar tree for any situation or data, we can limit the data to particular targets (e.g. kinases) or apply ligand-efficiency rules to the predictions. For more details see the talk above.
If you are interested in an evaluation version of the software used here, Matsy, send an email to matsy@nextmovesoftware.com.