There is a 2008 paper by the folks at Abbott that Roger refers to as “the paper that killed matched pairs for activity data”. This is the Hajduk and Sauer paper which looked at matched pair transformations across 84K compounds and 30 protein targets, and found that the potency changes associated with most matched pairs transformations were (nearly) normally distributed around zero.
I don’t have access to the Abbvie data, but I can do a similar analysis with ChEMBL data. For those assays in ChEMBL which have pIC50 data for ethyl, propyl and butyl as substituents at the same location in three molecules, here is a histogram of the pIC50 for the ethyl analog minus that of the butyl.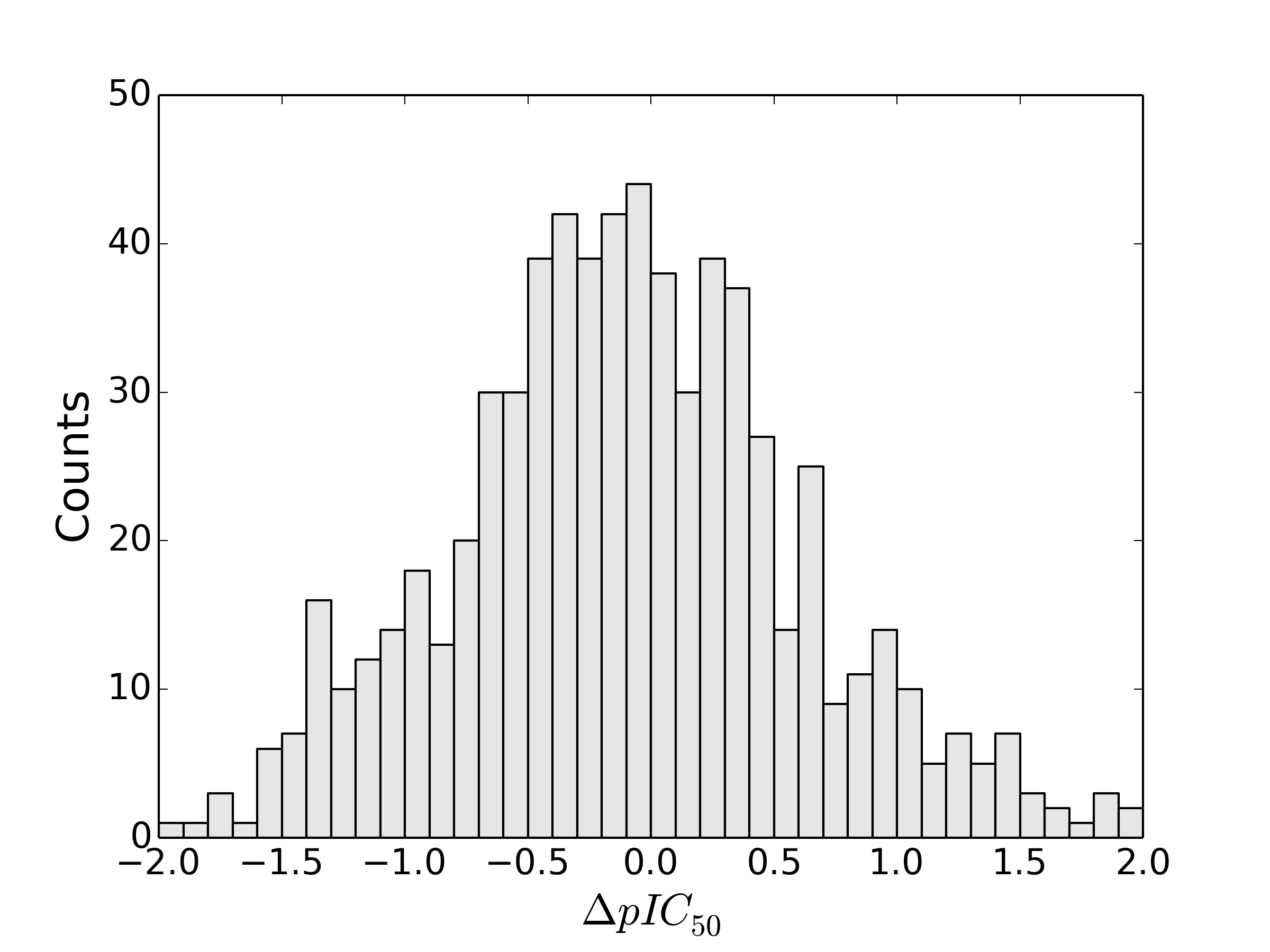
The resulting histogram is pretty much in agreement with what Hajduk found: changing a butyl to an ethyl is equally likely to increase activity as decrease it. If we think about it, it’s fairly obvious why this might be – we are pooling data from different binding environments in different proteins, and the effect of the change depends on the binding environment.
So matched pairs are dead for activity. Or we have to restrict the analysis to data just for a particular pocket in a particular protein.
But what if we consider additional activity data? What if we already know the relative activities of propyl and butyl, for example. Let’s say that we know in a particular case that the propyl analog has a greater pIC50 than butyl, and then plot the ΔpIC50 for ethyl minus butyl:
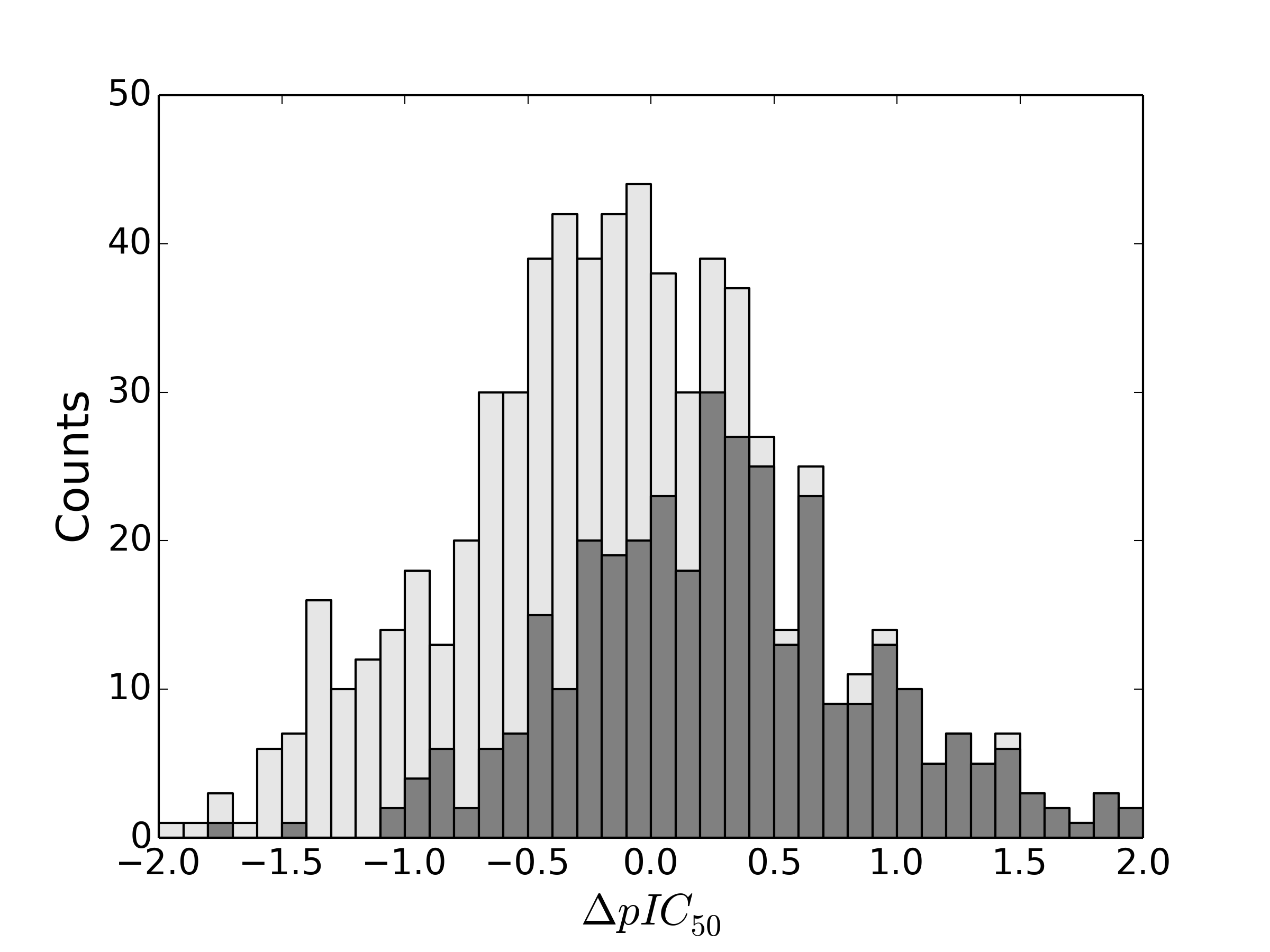
…or vice versa, where propyl has a smaller pIC50 than butyl:

Interesting, eh? The point is that knowing some activity information improves our predictive ability. If we know that propyl > butyl, then it increases the chance that changing butyl to ethyl will increase the activity.
The question then arises, how best to extract and apply this information? One approach would be to throw more matched pairs at the problem. But actually, a simpler and more elegant approach is to look beyond matched pairs to the general concept of matched series.
Here are the slides I presented on “Evidence-based medicinal chemistry with matched series” at the recent UK-QSAR meeting in Cambridge.